Project Goals
 I am a plant biologist with broad interests in processes and patterns of evolution at both the molecular and organismal levels. With the advent of genome sequencing, much of our work has centered on the generation and analysis of large scale sequence and expression datasets using bioinformatic and molecular evolutionary approaches. I am fortunate to maintain both wet-lab and dry-lab (informatic) components in my laboratory and to be able to welcome lab members with either or both research interests.
I am a plant biologist with broad interests in processes and patterns of evolution at both the molecular and organismal levels. With the advent of genome sequencing, much of our work has centered on the generation and analysis of large scale sequence and expression datasets using bioinformatic and molecular evolutionary approaches. I am fortunate to maintain both wet-lab and dry-lab (informatic) components in my laboratory and to be able to welcome lab members with either or both research interests.
My research focuses on three project areas:
- The evolution of the flower and the floral developmental program
- The study of parasitic plants
- Chloroplast genome and phylogenomic evolution
Evolutionary Genomics of the Flower

The sudden appearance of flowers in the fossil record over 100 million years ago represents a great mystery that has long puzzled plant and evolutionary biologists. Among the many key questions surrounding the origin and diversification of flowers include: Did the earliest flowers possess the full complement of genetic information needed to assemble a modern flower? Was genome duplication an important driver of the diversification of floral developmental pathways? Are there genes that have conserved functions throughout much of flowering plant history? Recent studies in plant developmental genetics and genomics have identified more than 100 genes with specific roles in flower development in Arabidopsis and other model organisms. Other genes with critical roles may remain undiscovered, largely because of functional redundancy or lethality of loss-of-function mutations. This knowledge, along with the recently clarified phylogenetic relationships of flowering plants (Leebens-Mack et al. 2005) and the availability of genome-scale data and bioinformatics analysis now make it possible to begin to understand how floral developmental pathways originated and diversified.
The Floral Genome Project, a mulit-institutional, multi-collaborator study funded through NSF's Plant Genome Research Project, was initiated to address these questions by studying the evolutionary diversification of floral regulatory genes and pathways throughout the major lineages of flowering plants. The study was designed to capture a large number of genes expressed during early flower development in 15 phylogenetically-critical lineages of flowering plants and gymnosperms, and to determine their expression patterns at several levels of resolution. We then link the sequences and expression patterns through phylogenetic and molecular evolutionary analysis to infer the gene sets and expression patterns that may have been present in the earliest angiosperm lineages. As the PI of the FGP, my group is involved in many aspects of the project, including library building, screening, and sequencing, studies of individual floral gene families, database construction, and a wide range of bioinformatic and molecular evolutionary studies. Read more...
Parasitic Plants

Another long term focus of our research is the study of parasitic plants -- phylogeny, biology, and molecular evolution. Although most plants are autotrophic, several thousand species of angiosperms obtain water, minerals, and fixed carbon heterotrophically, using specially modified roots (haustoria) that extract these materials directly from a host plant. In addition to their intrinsic interest as organisms with complex adaptations for direct feeding upon other plants, the hosts of some parasitic plants are important crop plants, and thus of great economic significance. Furthermore, because some parasites have completely lost the ability to photosynthesize, these plants provide a powerful system for the investigation of the effects of drastically altered functional constraints on gene and genome evolution and function. Read more...
Chloroplast Genomes
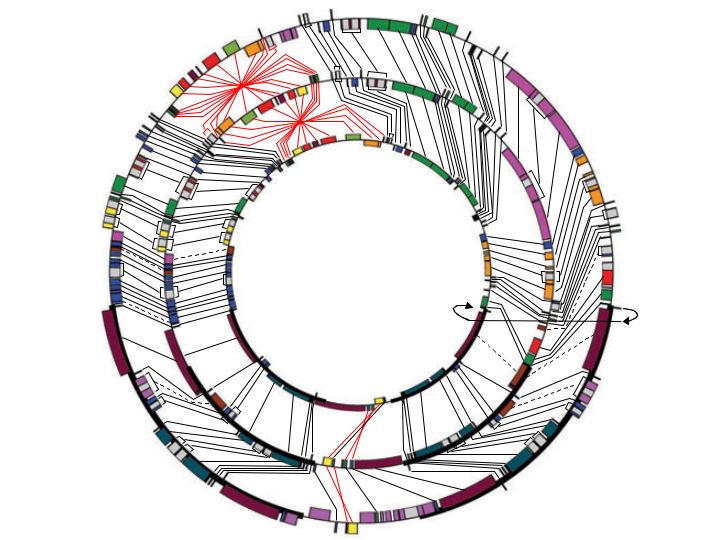
In plant cells, organelles called plastids are responsible for diverse functions, including photosynthesis, starch synthesis and storage, and key steps in lipid synthesis and nitrogen metabolism. Plastids are developmentally related to each other, with chloroplasts being the chlorophyll-containing plastids that perform photosynthesis. Many lines of evidence have shown that plastids are derived from endosymbiotic bacteria and contain DNA that is a very reduced genome of the bacterial endosymbiont. Most land plant chloroplast DNAs (cpDNAs) are similar, containing about 100-120 genes and possessing common structural features such as a large inverted repeat that contains the ribosomal rRNA genes. Although the first chloroplast genome sequence was published in 1986, the number of sequenced chloroplast genomes has remained limited until quite recently now that a number of novel cloning and sequencing strategies have been developed (see Jansen et al., 2005; McNeal et al., 2006). As a result, chloroplast genome sequencing has become much more routine and much less expensive. Through a project funded by NSF's Biodiversity and A Tree of Life (ATOL) programs, we are participating in the first large scale chloroplast genome sequencing project focused on angiosperms and other seed plants. Our goal is to sequence, annotate, and analyze more than 55 chloroplast genomes that represent all major lineages of flowering plants and gymnosperms, with a special focus on evolutionarily interesting genomes that have undergone rearrangements or gene content changes. Within this larger study, our lab's focus has been on the chloroplast genome evolution in parasitic plants, monocots, and basal angiosperms. Please see our chloroplast genome database and our publications page for recent papers stemming from this project.
Interested in doing research in our lab?
Please email me and let me know about your background and research interests. I am a faculty member in the Biology Department and the Huck Institutes of the Life Sciences at Penn State University, and participate in four graduate programs (see list below). From time to time, I can accept new undergraduates, graduate students, or postdocs, and I am always looking for highly motivated and well qualified individuals. So long as there is a good fit with the lab's interests, students can focus anywhere on the spectrum from organismal/developmental to pure bioinformatics. Graduate students should contact me to help identify the best program for your particular situation. Our doors are open to all nationalities and qualified minority applicants are especially encouraged.
Teaching
The classes I teach regularly include: Bioinformatics (Biol/Stat/CSE 597/598 teacher, taught with Webb Miller), Populations and Communities (Biol 220W, team taught with Jim Minesky; I teach the genetics component) and Taxonomy of Seed Plants (Biol 414). In addition, I usually contribute a few lectures to Genomics (IBIOS 598B) and Plant Communication and Growth Regulation (Plant Physiology 513). View course descriptions online at: www.bio.psu.edu/home/courses